matchedFilter-based peak detection in purely chromatographic data
Source:R/methods-Chromatogram.R
findChromPeaks-Chromatogram-MatchedFilter.RdfindChromPeaks on a MSnbase::Chromatogram() or
MSnbase::MChromatograms() object with a
MatchedFilterParam parameter object performs matchedFilter-based peak
detection on purely chromatographic data. See matchedFilter for details
on the method and MatchedFilterParam for details on the parameter class.
Note that not all settings from the MatchedFilterParam will be used.
See peaksWithMatchedFilter() for the arguments used for peak detection
on purely chromatographic data.
Usage
# S4 method for class 'Chromatogram,MatchedFilterParam'
findChromPeaks(object, param, ...)Arguments
- object
a
MSnbase::Chromatogram()orMSnbase::MChromatograms()object.- param
a MatchedFilterParam object specifying the settings for the peak detection. See
peaksWithMatchedFilter()for the description of arguments used for peak detection.- ...
currently ignored.
Value
If called on a Chromatogram object, the method returns a matrix with
the identified peaks. Columns "mz", "mzmin" and "mzmax" in
the chromPeaks() peak matrix provide the mean m/z and the maximum and
minimum m/z value of the Chromatogram object. See
peaksWithMatchedFilter() for details on the remaining columns.
See also
peaksWithMatchedFilter() for the downstream function and
matchedFilter for details on the method.
Examples
## Loading a test data set with identified chromatographic peaks
faahko_sub <- loadXcmsData("faahko_sub2")
faahko_sub <- filterRt(faahko_sub, c(2500, 3700))
#> Filter spectra
##
od <- as(filterFile(faahko_sub, 1L), "MsExperiment")
## Extract chromatographic data for a small m/z range
chr <- chromatogram(od, mz = c(272.1, 272.3))[1, 1]
## Identify peaks with default settings
xchr <- findChromPeaks(chr, MatchedFilterParam())
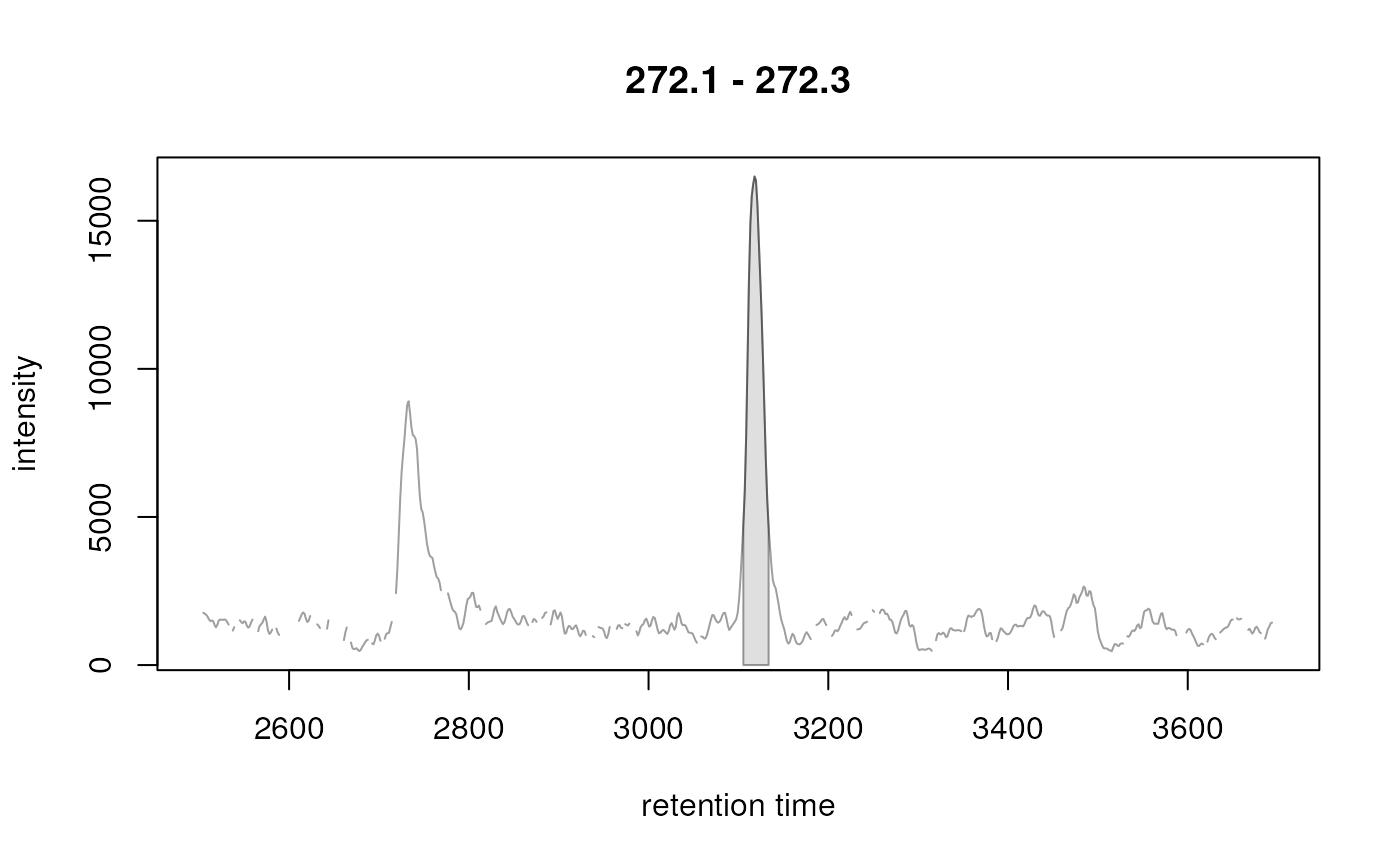
## Plot the identified peaks
plot(xchr)