
Correct gaps in data
stitch-methods.RdFixes gaps in data due to calibration scans or lock mass. Automatically detects file type and calls the relevant method. The mzXML file keeps the data the same length in time but overwrites the lock mass scans. The netCDF version adds the scans back into the data thereby increasing the length of the data and correcting for the unseen gap.
Methods
- object = "xcmsRaw"
stitch(object, lockMass=numeric())
- object = "xcmsRaw"
makeacqNum(object, freq=numeric(), start=1)
Arguments
- object
An
xcmsRaw-classobject- lockMass
A dataframe of locations of the gaps
- freq
The intervals of the lock mass scans
- start
The starting lock mass scan location, default is 1
Details
makeacqNum takes locates the gap using the starting lock mass scan and it's intervals. This data frame is then used in
stitch to correct for the gap caused by the lock mass. Correction works by using scans from either side of the gap to fill it in.
Value
stitch A corrected xcmsRaw-class object
makeacqNum A numeric vector of scan locations corresponding to lock Mass scans
Author
Paul Benton, hpaul.benton08@imperial.ac.uk
Examples
if (FALSE) library(xcms)
library(faahKO)
## These files do not have this problem to correct for but just
## for an example
cdfpath <- system.file("cdf", package = "faahKO")
cdffiles <- list.files(cdfpath, recursive = TRUE, full.names = TRUE)
xr<-xcmsRaw(cdffiles[1])
#> Create profile matrix with method 'bin' and step 1 ...
#> OK
xr
#> An "xcmsRaw" object with 1278 mass spectra
#>
#> Time range: 2501.4-4499.8 seconds (41.7-75 minutes)
#> Mass range: 200-600 m/z
#> Intensity range: 70-1373180
#>
#> MSn data on 0 mass(es)
#> with 0 MSn spectra
#> Profile method: bin
#> Profile step: 1 m/z (401 grid points from 200 to 600 m/z)
#>
#> Memory usage: 10.8 MB
##Lets assume that the lockmass starts at 1 and is every 100 scans
lockMass<-xcms:::makeacqNum(xr, freq=100, start=1)
## these are equcal
lockmass<-AutoLockMass(xr)
#> Warning:
#> Lock mass frequency wasn't detected
ob<-stitch(xr, lockMass)
ob
#> An "xcmsRaw" object with 1304 mass spectra
#>
#> Time range: 1.6-2040.7 seconds (0-34 minutes)
#> Mass range: 200-600 m/z
#> Intensity range: 70-1373180
#>
#> MSn data on 0 mass(es)
#> with 0 MSn spectra
#> Profile method: bin
#> Profile step: no profile data
#>
#> Memory usage: 6.96 MB
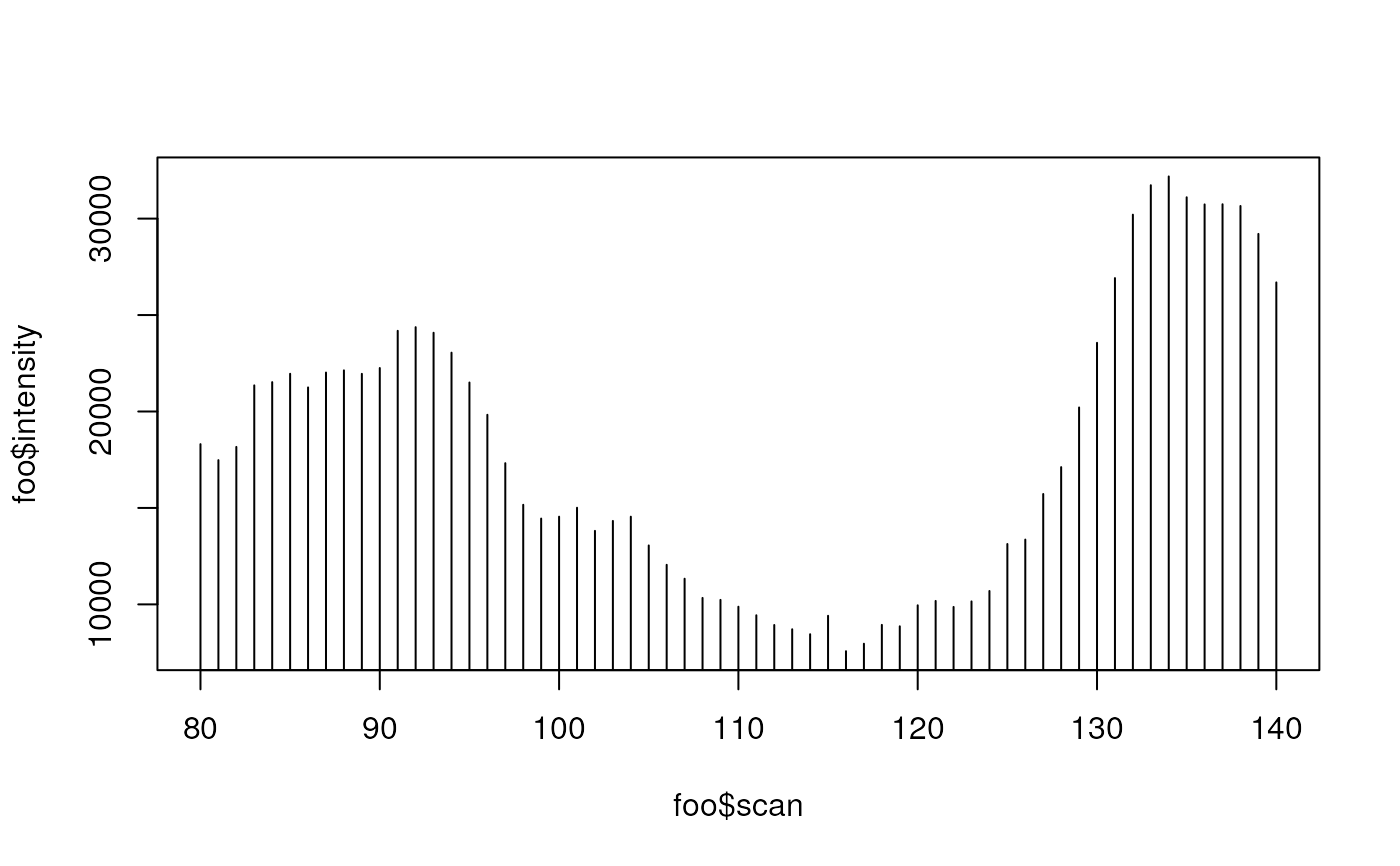
## plot the old data before correction
foo<-rawEIC(xr, m=c(200,210), scan=c(80,140))
plot(foo$scan, foo$intensity, type="h")
 ## plot the new corrected data to see what changed
foo<-rawEIC(ob, m=c(200,210), scan=c(80,140))
plot(foo$scan, foo$intensity, type="h")
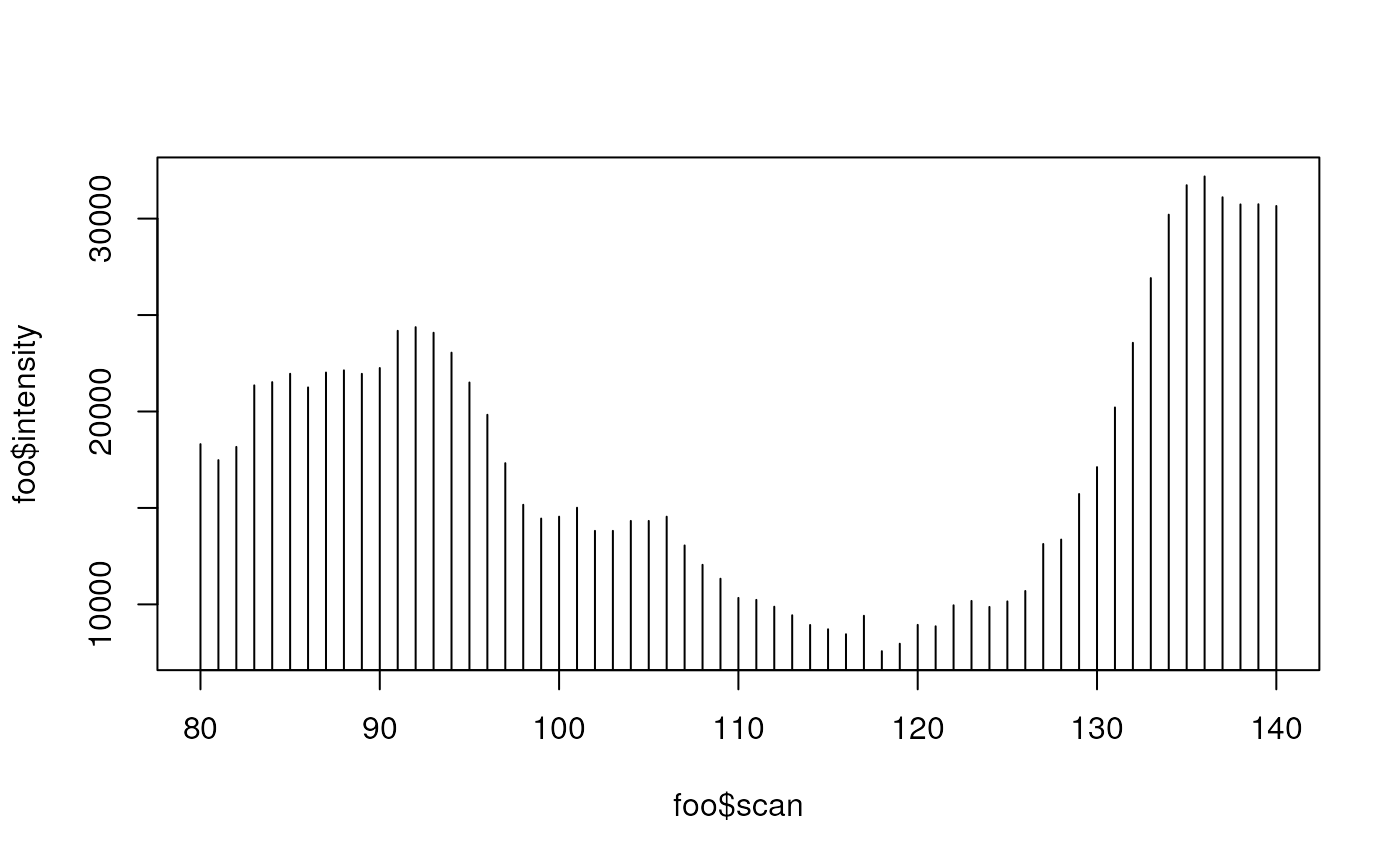
## plot the new corrected data to see what changed
foo<-rawEIC(ob, m=c(200,210), scan=c(80,140))
plot(foo$scan, foo$intensity, type="h")
 # \dontrun{}
# \dontrun{}